Tom Judd <<<>>> Math
For Compass Calibrations
Practical, Simplified routines applied to a Direct Least Squares Ellipse Solution
- ▶ Invert : Simple matrix inversion routine
- ▶ Eigenvalue/Eigenvectors : Eigenvalue/Eigenvectors of a Symmetric Matrix
- ▶ Cholesky : Robust Cholesky decomposition
- ▶ Direct Least Squares Ellipse : Robust implementation of Fitzgibbon, Pilu, and Fisher Direct Least Squares Fitting of Ellipses
Least Squares Fit to 2D and 3D Compass Data
- ▶ Circle: Quick, easy solution to simple 2D hard iron problem
- ▶ Sphere: Quick, easy solution to simple 3D hard iron problem
- ▶ Ellipse: 2D Hard and Soft iron solution
- ▶ Ellipsoid: 3D Hard and Soft iron solution
- ▶ Precision Ellipsoid: Precision Iterative 3D Hard and Soft iron solution
Invert
Most solutions for compass calibrations make use of a matrix inversion routine. Fortunately, these matrices are well behaved provided you have a good set of data. You have to have good data in order to calibrate a precision compass. If it blows up on you then it is a good indication that you have bad data.
I first ran into matrix inversion techniques in a numerical analysis course given in FORTRAN about 50 years ago. In the context of solving a set of linear equations one operates on a matrix using Gaussian elimination until it is in lower triangular form, then with back substitution, starting at the bottom, you work your way up, always having one equation and one unknown. Well, I didn't want to write any back substitution code, so I just continued zeroing out the upper triangle too. If you apply the same operations to an identity matrix, then the product of all those operations is the inverse. Magic!
Most matrix inversion routines want you to pivot, i.e. swap rows, so you don't divide by a small number. I dispense with that here with the expectation that all our diagonal elements are of reasonable size
With no pivoting, and no back-substitution, the code gets really simple. The end product consists of two matrices: the original matrix which has been reduced to an identity matrix, and the inversion matrix, which started out as an identity matrix. You can probably write it up yourself in C without peeking once you have read through this once. So let's start.
The sample matrix we start with is
[1.1 ,0.1, 0.05],
[0.1 ,1.2, 0.2 ],
[0.05 ,0.2, 1.3 ]
Shown in augmented form with the identity matrix appended to the end it is
1.100 0.100 0.050 1.000 0.000 0.000
0.100 1.200 0.200 0.000 1.000 0.000
0.050 0.200 1.300 0.000 0.000 1.000
We first normalize the first diagonal element (divide the row by the diagonal element)
1.0000 0.0909 0.0455
0.1000 1.2000 0.2000
0.0500 0.2000 1.3000
In the code, the part that does the normalization is shown below. Notice that we check
for zero before dividing. The code applies the division to all elements in the row AND
all elements of the augmented portion of the matrix. Some algorithms work on a single
augmented matrix and span 2N elements. I think this way is clearer.
alpha = A[ii][ii]
if abs(alpha) < epsilon: return (Ainv,-1)
for jj in range(N):
A[ii][jj] = A[ii][jj]/alpha
Ainv[ii][jj] = Ainv[ii][jj]/alpha
We then add a scaled version of it to the first element we want to eliminate. Since
it is 0.1 we multiply the first row by -0.1 and add it to the second row. We get
1.0000 0.0909 0.0455
0.0000 1.1909 0.1955
0.0500 0.2000 1.3000
The augmented part, which started out as an identity matrix, now reads
0.9091 0.0000 0.0000
-0.0909 1.0000 0.0000
0.0000 0.0000 1.0000
In the code, the part that does the elimination is shown below.
beta = A[kk][ii]
for jj in range(N):
A[kk][jj]=A[kk][jj] - beta*A[ii][jj]
Ainv[kk][jj]=Ainv[kk][jj] - beta*Ainv[ii][jj]
That's all there is to it. Just put it in a loop so that it iterates over all diagonal
elements (ii index). Then eliminate each element in the column (kk index) except itself
(if kk==ii: continue). The complete algorithm is in the code section.
Python Source Code mat_invert.py
import numpy as np
def mat_invert(A,epsilon=1E-10):
N = len(A)
Ainv=np.eye(N)
for ii in range(N):
alpha = A[ii][ii]
if abs(alpha) < epsilon: return (Ainv,-1)
for jj in range(N):
A[ii][jj] = A[ii][jj]/alpha
Ainv[ii][jj] = Ainv[ii][jj]/alpha
for kk in range(N):
if kk==ii: continue
beta = A[kk][ii]
for jj in range(N):
A[kk][jj]=A[kk][jj] - beta * A[ii][jj]
Ainv[kk][jj]=Ainv[kk][jj] - beta * Ainv[ii][jj]
return (Ainv,1)
if __name__ == "__main__":
from MUTIL import * # printMat and printVec utilities
import copy
C=np.array( [ [1.1,0.1,0.05],
[0.1,1.2,0.2 ],
[0.05,0.2,1.3] ])
svC=copy.deepcopy(C)
printMat(C,"%12.6f",'C mat')
invC,ok=mat_invert(C)
printMat(invC,"%12.6f",'C invert')
invCC=np.dot(invC,svC)
printMat(invCC,"%12.6f",'C invert times C')
**********Sample Output*********
C:\SCRIPTS\PYTHON\MATH>python mat_invert.py
C mat ( 3 , 3 )
1.100000 0.100000 0.050000
0.100000 1.200000 0.200000
0.050000 0.200000 1.300000
C invert ( 3 , 3 )
0.916767 -0.072376 -0.024125
-0.072376 0.860977 -0.129674
-0.024125 -0.129674 0.790109
C invert times C ( 3 , 3 )
1.000000 0.000000 0.000000
-0.000000 1.000000 0.000000
-0.000000 0.000000 1.000000
Eigenvalues/Eigenvectors
Most compass calibrations algorithms invert a matrix to find the coefficients of the polynomial for an ellipse or ellipsoid. The solution is then shifted to the center, and an eigenvalue / eigenvector routine is applied in order to determine the full transformation matrix of the solution.
In some algorithms, the polynomials are also obtained using an eigenvalue / eigenvector routine. So, it is important to look at an eigenvalue routine.
A simple eigenvalue routine was derived by Kaiser (see references). I ran across it 40 years ago when looking for a method that was faster than the Givens-Householder method. It wasn't. But it is simpler. So, I cover it here.
I modified the routine slightly to address some of the problems encountered in solving for the polynomials using the direct least squares Fitzgibbon algorithm. The original routine was designed for matrices with positive eigenvalues. The Fitzgibbon method relies on a matrix that can have eigenvalues that are zero or negative. I found that the JK method worked for negative values, but since we have to divide by the eigenvalues to normalize the vectors, I eliminated solutions that were too close to zero. A passed parameter sets the zero threshold:
#set vectors to zero whose eigenvalues are too close to zero
absV=abs(values[jj])
if absV < Zthreshold:
for ii in range(N): vectors[ii][jj]=0
else:
for ii in range(N): vectors[ii][jj]=A[ii][jj]/values[jj]
If your matrices have all positive eigenvalues, then you can skip the step that returns the negative values. See the comments in the code.
The B matrix in the sample data has three eigenvalues that are too close to zero. See the output at the bottom of the listing. For comparison, I used the numpy library eig function. This matrix is an example of one that might be found using the direct least squares algorithm. Even though we get multiple eigenvalues / eigenvectors, only the eigenvector associated with the largest positive eigenvalue is used for the solution.
If all this seems too complicated, see Tom's Shortcut method that completely eliminates the need for an eigenvalue / eigenvector routine. Only a simple matrix inversion routine is required. (see above). This is ultra-handy in embedded systems where it might not be feasible to include a general eigenvector routine that can also solve for asymmetric matrices.
Python Source Code JK.py
# Compute the eigenvalues and eigenvectors for a symmetric matrix.
# Matrix is real. Eigenvectors are normalized with (divided by) the eigenvalue.
# For positive definite matricies this is not a problem. Otherwise, compare the
# eigenvalues to a threshold, and if too close to zero, reject the solution.
# From:
# The JK Method: A procedure for finding the eigenvalues of a real
# symmetric matrix, H.F.Kaiser, The Computer Journal, Vol. 15 (1972), pp 271-273
import sys
import numpy as np
import copy
# Equation numbers are from original paper
# The JK method: a procedure for finding the eigenvectors
# and eigenvalues of a real symmetric matrix.
# by H. F. Kaiser
def JK(A,Zthreshold=1e-6):
epsilon=1.0e-16
converge=1.0e-8
verbose=0
N=len(A)
values=np.zeros(N)
vectors=np.zeros((N,N))
lastDiag=0.0
svA=copy.deepcopy(A)
for iterLoop in range(15):
for jj in range(N-1):
for kk in range(jj+1,N):
num=0.0 #in the FORTRAN code, num is 'P' and den is 'Q'
den=0.0
for ii in range(N):
num = num + A[ii][jj] * A[ii][kk] # numerator eq. 11
den = den + (A[ii][jj] + A[ii][kk]) * (A[ii][jj] - A[ii][kk]) # denominator eq. 11
num = 2*num
anum=abs(num)
aden=abs(den)
if anum < epsilon and den >=0:
break # halt k loop
if anum <= aden:
tan2 = anum / aden #eq. 11
cos2 = 1 / np.sqrt(1 + tan2 * tan2) #eq. 12
sin2 = tan2 * cos2 #eq. 13
else:
cot2 = aden / anum #eq. 16
sin2 = 1 / np.sqrt(1 + cot2 * cot2) #eq. 17
cos2 = cot2 * sin2 #eq. 18
ca = np.sqrt((1 + cos2) / 2) #eq. 14/19
sa = sin2 / (2 * ca) #eq. 15/20
if den < 0 :
tmp = ca
ca = sa # table 21
sa = tmp
if num < 0 :
sa= -sa
for ii in range(N):
tmp = A[ii][jj]
A[ii][jj] = tmp * ca + A[ii][kk] * sa
A[ii][kk] = -tmp * sa + A[ii][kk] * ca
da=np.diagonal(A)
dadot=np.dot(da,da)
if verbose==1:
print dadot
if abs(dadot-lastDiag) < converge:
if iterLoop > 5:
break # stop iterations if diagonal is not changing
lastDiag=dadot
for jj in range(N):
for kk in range(N):
values[jj]=values[jj] + A[kk][jj]* A[kk][jj]
values[jj] = np.sqrt(values[jj]) # these will all be positive, OK for positive definite matrix
#set vectors to zero whose eigenvalues are too close to zero
absV=abs(values[jj])
if absV < Zthreshold:
for ii in range(N): vectors[ii][jj]=0
else:
for ii in range(N): vectors[ii][jj]=A[ii][jj]/values[jj]
# get signed (negative) eigenvalues for non-positive definite matrix
# you can skip this step and return 'values' instead of 'svalu'
# if you are working with only positive eigenvalues
svalu=np.diagonal(np.dot(vectors.T,np.dot(svA,vectors)))
return (svalu,vectors)
if __name__ == "__main__":
from numpy.linalg import eig, inv
def printVec( vec, sfmt="%9.4F", title="vec" ):
ii = len(vec)
print '\n',title,'(',ii,')'
ss = " "
for ix in range(ii):
ari=np.real(vec[ix])
sad = sfmt % ari
ss = ss + sad
print ss
return
from numpy.linalg import eig, inv
def printMat( arr, sfmt="%9.4f", title="mat" ):
ii = len(arr)
jj = len(arr[0])
print '\n',title,'(',ii,',',jj,')'
for ix in range(ii):
ari=arr[ix]
ss = " "
for jx in range (jj):
arj=ari[jx]
sad = sfmt % arj
ss = ss + sad
print ss
return
zeroThreshold = 1e-6
B=np.array([[ 0.000000, 0.000000, 0.055735, -0.182271, -1.362358, 1.544174],
[ 0.000000, -0.029843, 0.016174, -0.047248, 2.517891, -2.782243],
[ 0.055735, 0.016174, -0.112859, -2.046633, -15.374392, 32.096392],
[ -0.182271, -0.047248, -2.046633, 14.592118, 108.775746,-171.281981],
[ -1.362358, 2.517891, -15.374392, 108.775746, 534.644957,-971.581651],
[ 1.544174, -2.782243, 32.096392,-171.281981,-971.581651,1515.448501]])
C=np.array( [ [1.0,0.1,0.05],
[0.1,1.0,0.2 ],
[0.05,0.2,1.0] ])
#in place modification of B and C matrices, so we need to keep a reference copy around
svC=copy.deepcopy(C)
svB=copy.deepcopy(B)
printMat(B,"%12.6f",'B mat')
# conventional eigenvalue/eigenvectors from numpy library
valB,vecB=eig(B)
vv=np.real(vecB)
print "\nSolution using numpy library"
printVec(valB,"%12.5f","B EigenValues")
printMat(vecB,"%12.6f",'B Eigenvectors')
# solution using JK algorithm
(valBjk,vecBjk)=JK(B,zeroThreshold)
vecBjk=np.real(vecBjk)
printVec(valBjk,"%12.5f","JK B EigenValues")
printMat(vecBjk,"%12.6f",'JK B eigenvectors')
# do the same for a positive definite matrix
printMat(C,"%12.2f",'C mat')
valC,vecC=eig(C)
vv=np.real(vecC)
printVec(valC,"%12.5f","C numpy EigenValues")
printMat(vecC,"%12.6f",'C numpy Eigenvectors')
(valC,vecC)=JK(C,zeroThreshold)
printVec(valC,"%12.5f","C JK EigenValues")
printMat(vecC,"%12.6f",'C JK Eigenvectors')
*********SAMPLE OUTPUT**********
B mat ( 6 , 6 )
0.000000 0.000000 0.055735 -0.182271 -1.362358 1.544174
0.000000 -0.029843 0.016174 -0.047248 2.517891 -2.782243
0.055735 0.016174 -0.112859 -2.046633 -15.374392 32.096392
-0.182271 -0.047248 -2.046633 14.592118 108.775746 -171.281981
-1.362358 2.517891 -15.374392 108.775746 534.644957 -971.581651
1.544174 -2.782243 32.096392 -171.281981 -971.581651 1515.448501
Solution using numpy library
B EigenValues ( 6 )
2133.40657 -63.64643 -5.21727 0.00000 -0.00000 -0.00000
B Eigenvectors ( 6 , 6 )
-0.000955 -0.005524 0.000923 -0.474946 -0.708142 0.522426
0.001719 0.010975 0.073883 0.294578 -0.685592 -0.661517
-0.016602 0.058110 -0.305205 0.793816 -0.168265 0.494713
0.095320 0.047018 0.944175 0.231153 0.000111 0.209299
0.521741 -0.851007 -0.026389 0.048541 -0.012637 0.019002
-0.847597 -0.519662 0.096064 0.041459 -0.005063 0.023614
JK B EigenValues ( 6 )
2133.40657 -63.64643 -5.21727 0.00000 0.00000 0.00000
JK B eigenvectors ( 6 , 6 )
0.000955 0.005524 0.000923 0.000000 0.000000 0.000000
-0.001719 -0.010975 0.073883 0.000000 0.000000 0.000000
0.016602 -0.058110 -0.305205 0.000000 0.000000 0.000000
-0.095320 -0.047018 0.944175 0.000000 0.000000 0.000000
-0.521741 0.851007 -0.026389 0.000000 0.000000 0.000000
0.847597 0.519662 0.096064 0.000000 0.000000 0.000000
C mat ( 3 , 3 )
1.00 0.10 0.05
0.10 1.00 0.20
0.05 0.20 1.00
C numpy EigenValues ( 3 )
1.24622 0.96075 0.79303
C numpy Eigenvectors ( 3 , 3 )
0.399264 0.896252 0.193187
0.670284 -0.141577 -0.728474
0.625545 -0.420344 0.657270
C JK EigenValues ( 3 )
1.24622 0.96075 0.79303
C JK Eigenvectors ( 3 , 3 )
0.399264 -0.896252 0.193187
0.670284 0.141577 -0.728474
0.625545 0.420344 0.657270
Robust Cholesky Decomposition
A Cholesky decomposition of a matrix creates a 'square root' lower triangular matrix which when multiplied by its transpose recovers the original matrix:
A = L*LT
Standard algorithms iterate through each row and form at some point a divisor in the form of a difference:
d = A[ii][ii] -S
We can't let this divisor go zero or negative. With noisy data it can. We can rescue the algorithm by forcing suspicious divisors to be a small positive number. The definition of 'small' depends on the data. In this algorithm I look at the ratio of the smallest to the largest divisor to get an idea of the scale of the problem. If it is too tiny, I make the substitution.
The justification for this approach is that we know there is a real solution to the problem. Our data may be marginal in just the wrong places to trigger a zero or negative divisor. So, we fix the data. This adds a small amount of noise. But for compass calibrations, I have found that the noise is insignificant by orders of magnitude relative to the accuracy of the compass.
Python Source Code choleskyx.py
def choleskyX(A, ztol= 1.0e-5):
"""
Computes the lower triangular (LT) Cholesky factorization of
a positive definite matrix A.
"""
# Forward step for symmetric triangular L.
nrows = len(A)
errFlag=0
ztol2=ztol*ztol
L = np.zeros((nrows,nrows))
diag=np.diagonal(A)
absdmin=np.sqrt(np.dot(diag,diag))
absmax=1 # division by absmax below will always be defined
# print "diag ",absdmin
for ii in range(nrows):
S=0.0
for k in range(ii):
S=S + L[k][ii]**2
d = A[ii][ii] -S
absd=abs(d)
if absd < absdmin:
absdmin=absd
if absd > absmax:
absmax = absd
absratio = absdmin/absmax
# print "absmax absmin absratio",absmax,absdmin,absratio
if absratio < ztol2 or d<0:
L[ii][ii]=ztol*np.sqrt(absmax)
rat = absd/absmax
# print "*** ",d," replaced by ",L[i][i]
print "*** Tolerance: ",ztol, " greater then ",np.sqrt(rat)
errFlag = -1
else:
L[ii][ii] = sqrt(d)
for j in range(ii+1, nrows):
S=0.0
for k in range(ii):
S = S + L[k][ii]*L[k][j]
# if abs(S) < ztol:
# S = 0.0
L[ii][j] = (A[ii][j] - S)/L[ii][ii]
L=L.T
return(L,errFlag,absdmin)
Direct Least Squares Ellipse
Robust implementation of the Fitzgibbon, Pilu, and Fisher algorithm
The direct least squares method of Fitzgibbon, Pilu, and Fisher algorithm has two important features:
- It is simple to implement. The original article solves everything with seven lines of MATLAB code
- The constraint in the equations forces the solution to an ellipse (instead of another conic section). This is useful in situations where the data is noisy or marginal
The big drawback is that it is unstable with very low noise data. Consequently, Halir and Flusser developed a method of stabilization based on partitioning the matrix in question so that its eigenvalues were well behaved with noisy data.
Here, I develop a shorter and more pragmatic solution for compass calibrations. I split the matrix with a Cholesky decomposition. Unstable matrices will crash. I impose stability by forcing zero and small negative divisors in the Cholesky decomposition to be slightly positive. I have found that the errors introduced are smaller by multiple factors of ten compared to the compass calibration errors. Let's start from Fitzgibbon's eigenvalue equation:
S a = λ C a
a = λ S-1 C a [Eq. 1]
We want to find the eigenvalues and eigenvectors of S-1 C . There are two problems:
- The matrix is unstable
- The matrix is asymmetric
S = L LT
We also substitute
a = ( L-1 )T b [Eq. 2]
and use
( A B )T = BT AT and ( A-1 )T = ( AT )-1
Plugging all of this into equation 1 above we arrive at
( L-1 )T b = λ( L-1 )T L-1 C ( L-1 )T b
or
b = λ( L-1 ) C ( L-1 )T b [Eq. 3]
or restating as a simple eigenvalue equation:
b = λ(symmetric matrix) b
So, we just use our simple SYMMETRIC matrix algorithm to get the eigenvectors b, and then use equation 2 to get the original eigenvectors a.
Sample of Robust implementation of Direct Least Squares Ellipse Algorithm
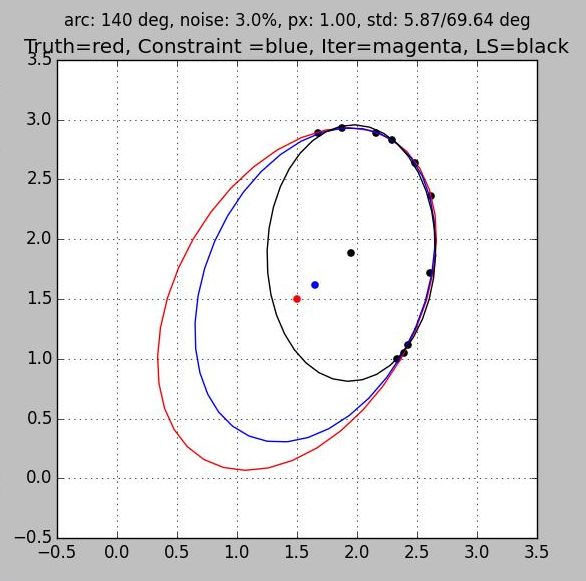
Python Source Code
The source code uses the three algorithms listed above:
- choleskyX.py for the robust Cholesky decomposition of the S matrix
- mat_invert.py to obtain the inverse of the Cholesky matrix
- jk.py to obtain the eigenvalues and eigenvectors of a symmetric matrix
The source shows how to obtain the S and C matrices from the input data, and then picks up with the text to the left:
- The robust Cholesky decomposition is performed on S
L,err,threshold=choleskyX(S,tolerance) - The inverse and transpose of L are calculated
(invL,status)=mat_invert(L) invLT = invL.T - The symmetric matrix of equation 3 CC is formed and its eigenvectors determined:
CC=np.dot(invL,np.dot(C,invLT)) valC,vecC=JKOBI.JK(CC) - Finally, the eigenvectors for the original equation 1 are obtained via equation 2:
vvcl=np.dot(invLT,vecC)
def ls_ellipse_robust(xin,yin,tolerance=1e-5,verbose=0):
x = xin[:,np.newaxis]
y = yin[:,np.newaxis]
D = np.hstack((x*x, x*y, y*y, x, y, np.ones_like(x)))
S = np.dot(D.T,D)
C = np.zeros([6,6])
C[0,2] = C[2,0] = 2; C[1,1] = -1
if verbose> 0:
print "tolerance is ",tolerance
printMat(D,"%12.6f", "np Dmat" )
printMat(S,"%12.6f", "np Smat" )
printMat(C,"%12.6f", "np Cmat" )
try:
#tolerance is used to check for division by zero
L,err,threshold=choleskyX(S,tolerance)
if verbose > 0:
printMat(L,"%12.6f", "L" )
if err < 0:
print "Min L Diagonal ",threshold
(invL,status)=mat_invert(L)
if status < 0:
print "Invert L error, near zero diagonal"
invLT = invL.T
CC=np.dot(invL,np.dot(C,invLT))
if verbose > 0:
printMat(invL,"%12.6f", "inverse L" )
printMat(CC,"%12.6f", "lClT (CC matrix being eigened)" )
valC,vecC=JKOBI.JK(CC)
vvcl=np.dot(invLT,vecC)
vvcl=np.real(vvcl)
#get eigenVector from index of largest eValue
ix=np.argmax(valC) # JK sorts vectors so this should be zero
if verbose > 0: print 'Index is ',ix
cholv=vvcl.T[ix][:]
nrm=np.sqrt(np.dot(cholv,cholv))
cnrm=cholv/nrm
if verbose> 0:
printVec(valC,"%12.5f","tj EigenValues")
printMat(vvcl,"%12.5f","tj EigenVectors")
printVec(cnrm,"%12.5f", "Return normalized polynomial : ")
except:
print "\nfit tj error"
return (-1,0,0,0)
return (1,cnrm,valC,vvcl)
©2022 Tom Judd
www.juddzone.com
Carlsbad, CA
USA